Engineering Precision Medicine: How IntegraMS Translates Omics Data into Actionable Clinical Insights
From Biology to Engineering
At its core, IntegraMS is a full-stack, machine learning–driven application that bridges high-dimensional proteomic data with clinical outcomes in Multiple Sclerosis (MS).The problem is deceptively complex: clinicians measure antibody responses (via phage display), serum neurofilament light chain (nFL), and EDSS disability scores at different timepoints, yet these rich data streams remain disconnected.The IntegraMS system unifies them through a reproducible cloud-based pipeline — integrating data ingestion, preprocessing, prediction, and visualization into one cohesive workflow.
1. Data Flow and Synthetic Data Generation
The foundation of the IntegraMS infrastructure is a hierarchical data model that captures patient and visit-level relationships.
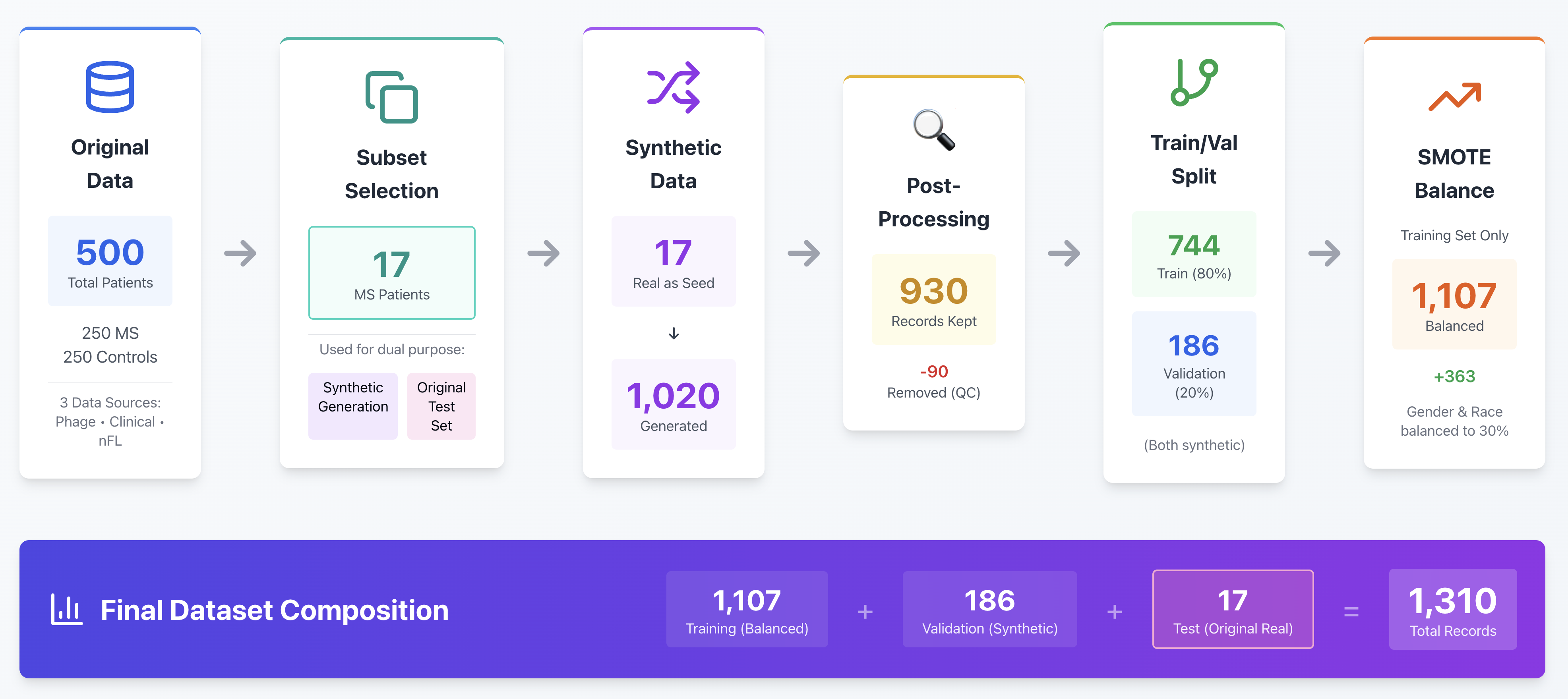
As illustrated below, real-world data (proteomic, clinical, and nFL) is first ingested, metadata is defined, and the Hierarchical Modeling Algorithm (HMA) Synthesizer generates synthetic analogs.
.png)
This Synthetic Data Vault (SDV) approach enables testing, bias mitigation, and reproducibility on small sample sizes (≈500 patients, ~400,000 records).Each patient record is modeled across timepoints — pre- and post-diagnosis — allowing synthetic replication that maintains the hierarchical structure of patient–visit relationships without compromising privacy
2. The Machine Learning Engine: One-Step MS Progression Model
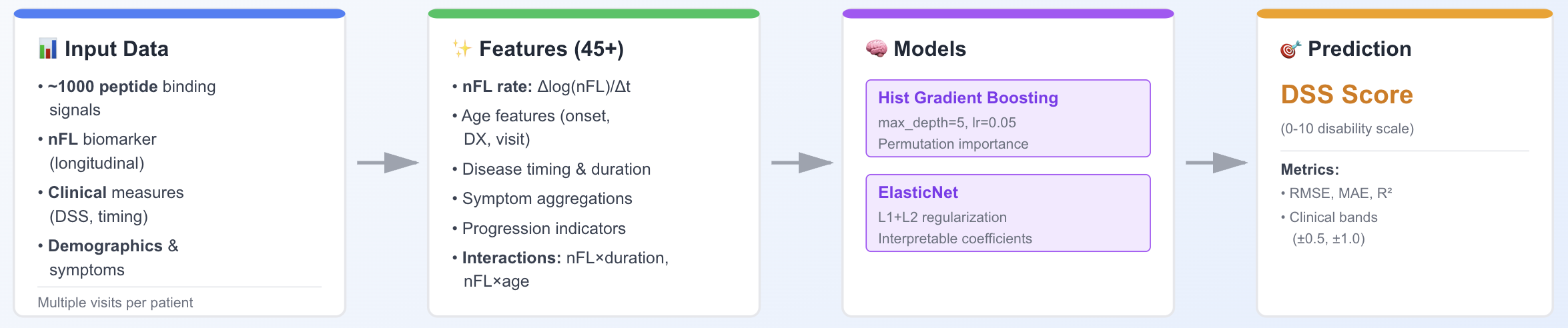
At the center of IntegraMS lies a supervised ML pipeline designed to model MS disability status directly. Rather than chaining multiple stages together, the system focuses on predicting the current Disability Status Scale score (DSS_Cur) on its 0–10 scale from a combination of curated proteomic features and aligned clinical metadata. The model ingests demographic variables, disease timeline features (such as time from onset to diagnosis and time since diagnosis), longitudinal DSS history, and a filtered set of peptide-level features derived from phage display assays. We train and evaluate regression models on a combined real and synthetic cohort using patient-level cross-validation, with metrics including MAE, RMSE, R², and a clinically motivated tolerance of ±1 DSS point. Feature importance and attribution analyses highlight the proteomic and clinical variables most associated with disability status, allowing us to connect model behavior back to emerging MS biomarker findings in the literature and to the underlying biology of disease progression.
3. Backend Architecture: Cloud-Native ML Inference
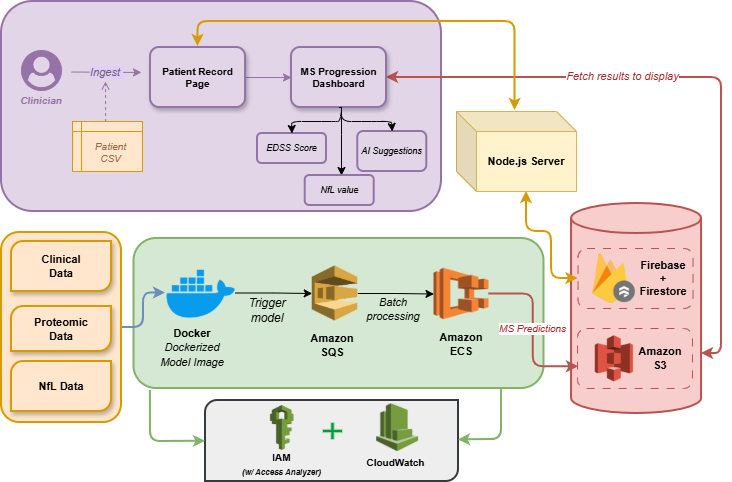
IntegraMS is deployed as a modular microservices architecture leveraging AWS, Firebase, and Dockerized ML models.Each ML model is packaged as a Docker image — ensuring version control, reproducibility, and scalable deployment.
Cloud monitoring and IAM-based access control are implemented using AWS CloudWatch and Identity & Access Management (IAM), ensuring secure, traceable operations.

- Data Ingestion: Patient CSVs are uploaded via a secure web form.
- Storage: Files and metadata are stored in AWS S3 (for structured data) and Firebase Firestore (for patient/session management).
- Preprocessing & Prediction: A FastAPI service invokes trained ML containers hosted on Amazon ECS, triggered via Amazon SQS queues.
- Response Delivery: Results are served back through a Node.js API to the frontend in JSON format for visualization.
4. Integrated AI and Real-Time Insights
The next layer integrates Amazon Bedrock and OpenAI LLM APIs for real-time text summarization, interpretability, and clinician-friendly feedback.
When an analysis is complete, the LLM contextualizes results into a narrative report — explaining predicted EDSS scores, nFL trends, and biomarker relevance.
These insights are then surfaced in the MS Progression Dashboard, built with React and Tailwind CSS, allowing clinicians to review AI explanations and visualize trends interactively.

5. Frontend Application: From CSV to Clinical Insight
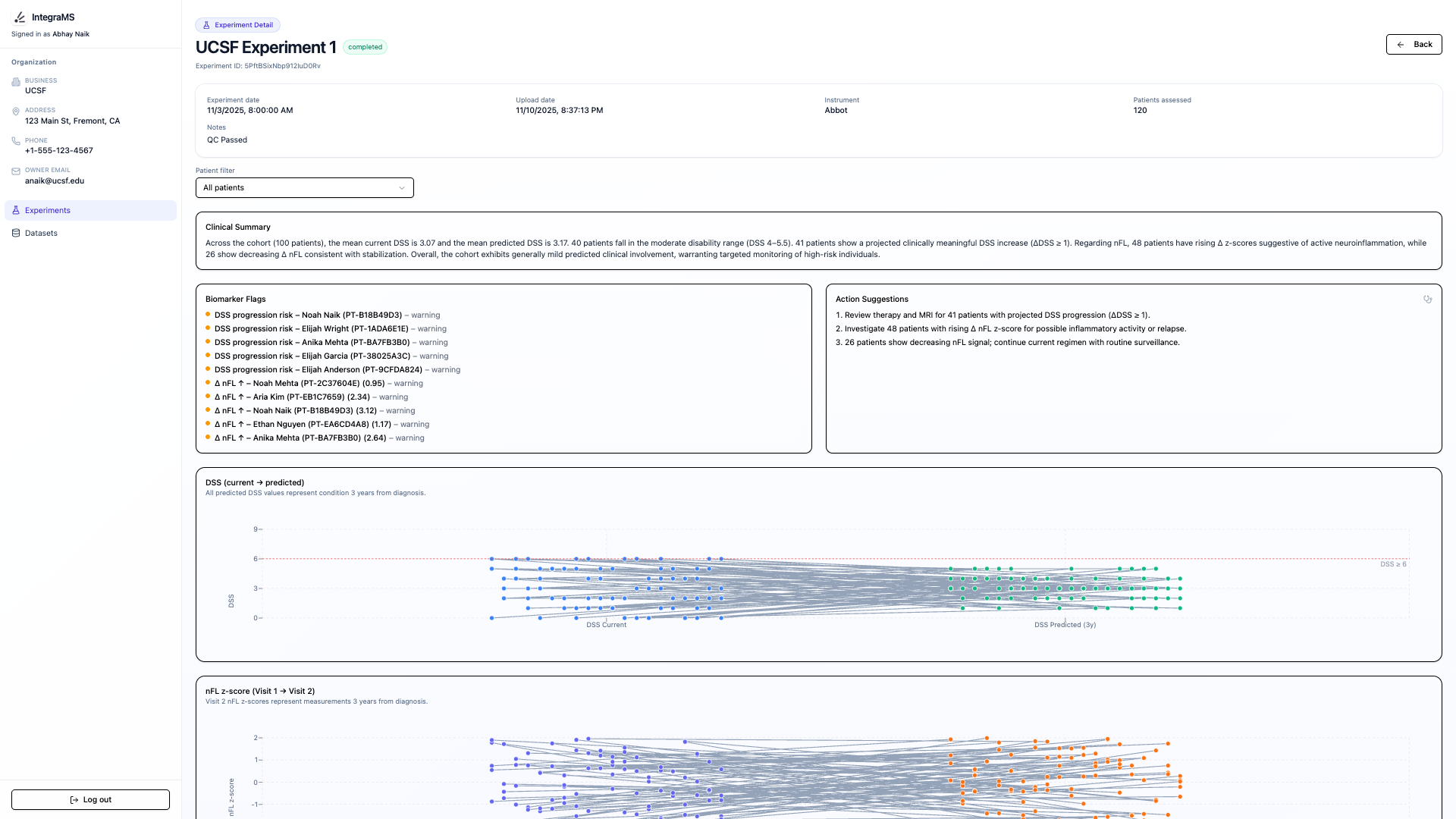
The frontend provides a seamless user experience for researchers and clinicians. Users can upload patient data in CSV format, trigger model runs, and visualize predicted DSS values alongside key clinical context. The dashboard surfaces disability predictions, timeline-aligned clinical information, and curated summaries of peptide-level features associated with each patient’s profile. Each result card is generated dynamically from the backend API and designed for clarity, interpretability, and auditability, reflecting the priorities and feedback gathered from clinicians during our primary research.
6. Security, Ethics, and Reproducibility
All patient-level data is de-identified before ingestion. The system adheres to HIPAA, GDPR, and FERPA standards with AES-256 encryption at rest and TLS 1.2 in transit.
Bias mitigation strategies include demographic stratification and synthetic data augmentation using SDV.
Clinician trust and explainability remain central — models are audited regularly for drift and interpretability consistency, aligning with the reproducibility principles outlined by Emmert-Streib (2022) and Mohr et al. (2024).
Conclusion: Building Scalable, Transparent AI for Precision Neurology
IntegraMS demonstrates how modern cloud infrastructure and ML engineering can translate raw omics data into clinically interpretable predictions.
By combining React-based visualization, FastAPI microservices, AWS orchestration, and LLM-driven interpretability, the system delivers both technical robustness and clinical usability.
IntegraMS stands as a reproducible, transparent, and clinically aware ML platform - bridging the gap between biological complexity and precision patient care.


